 
SLiMs are first presented in a table and when activated all instances of that SLiM are highlighted, thus displaying the distribution of the SLiM. 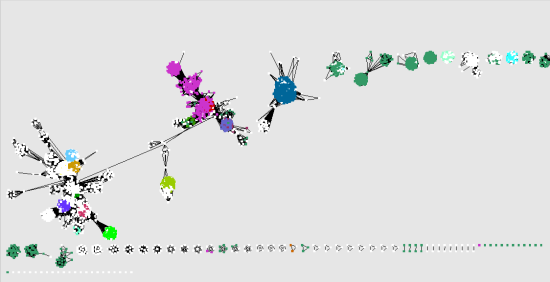
SLiMScape presents an interactive view of results within a protein interaction network. This functionality is implemented using SLiMSearch. Disorder and conservation masking are used to reduce the number of false positives. This is especially useful when trying to find new instances of a SLiM found using SLiMFinder and can also be used to search for a user defined SLiM. New instances of a motif can be located using regular expression matching. Protein sets can also be created manually, allowing for many custom strategies for finding motifs. This is useful when searching for self interacting proteins and other sequences which bind to a domain. The “Batch Domains” selection method creates sets of proteins which contain a particular domain, while “Batch Domains Interactions” creates sets of proteins which interact with a particular domain. A useful example is subcellular location, which can be used to find subcellular targeting motifs. Another selection method named “Attribute” creates protein sets based on a user-defined attribute. Since, in a large network, this can be computationally demanding selection methods include carrying out a single search across a set of selected nodes, specified interactively through Cytoscape’s interface. Figure 1 shows the results of this method on a network consisting of Proliferating Cell Nuclear Antigen (PCNA) and its interactors. This will iteratively create protein sets containing each protein (hub) and its interactors (spokes). For example, SLiM-mediated interactions can be discovered using the “Batch Interactions” selection method. A variety of node selection methods are provided to automatically select protein sets and different types of motif can be targeted depending on this selection process. SLiMScape uses SLiMFinder to locate statistically over-represented sequences within sets of proteins, defined by selecting subsets of a protein interaction network. An important feature is the visualization of results within a protein interaction network and in the context of domains in nearby proteins. The second, SLiMSearch can find new instances of known or potential motifs within the same network. SLiMFinder has previously been demonstrated as a suitable tool for mining the human interactome for SLiM mediated interactions. The first, SLiMFinder, is capable of finding novel motifs by analyzing a protein interaction network. SLiMscape consists of two primary analysis tools. Cytoprophet, another Cytoscape plugin, predicts domain and protein interaction networks however SLiMScape is the first to focus on SLiMs. Both novel and known motifs can be detected, providing a powerful tool for exploration and analysis of SLiMs within a protein interaction network. We have developed SLiMScape, a plugin which uses two of these tools to add SLiM discovery and search functionality to Cytoscape. Several predictive tools using many different methods have been developed to identify SLiMs. These often occur in intrinsically disordered regions and are involved in a number of functions such as binding, cleavage, subcellular targeting and post translational modifications. Many of these interactions occur between large globular domains but an estimated 15 - 40% are mediated by functional microdomains, of 3 - 10 amino acids in size called Short Linear Motifs (SLiMs). However, most of these experiments do not indicate the mechanism of interaction. High throughput experiments have greatly increased the number of known protein-protein interactions in the human interactome. This significantly aids in the discovery of novel short linear motifs and in visualizing the distribution of known motifs. SLiMScape provides a platform for performing short linear motif analyses of protein interaction networks by integrating motif discovery and search tools in a network visualization environment. 
To facilitate this SLiMScape automatically retrieves domains for each protein. The distribution of discovered or user-defined motifs may be selectively displayed and the distribution of protein domains may be viewed simultaneously. Data is presented using Cytoscape’s visualization features thus providing an intuitive interface for interpreting results. We present SLiMScape, a Cytoscape plugin, which enables both de novo motif discovery and searches for instances of known motifs. Cytoscape provides a visual interface for protein networks but there is no streamlined way to rapidly visualize motifs in a network of proteins, or to integrate computational discovery with such visualizations. Computational protein short linear motif discovery can use protein interaction information to search for motifs among proteins which share a common interactor. 
0 Comments
Leave a Reply. |
AuthorWrite something about yourself. No need to be fancy, just an overview. ArchivesCategories |
 RSS Feed
RSS Feed
